Similar to other predict methods, this functions predicts fitted
values, coefficients and more from a fitted ggmix_fit object.
# S3 method for ggmix_fit predict( object, newx, s = NULL, type = c("link", "response", "coefficients", "all", "nonzero", "individual"), covariance, ... ) # S3 method for ggmix_fit coef(object, s = NULL, type, ...)
Arguments
| object | Fitted |
|---|---|
| newx | matrix of values for |
| s | Value(s) of the penalty parameter |
| type | Type of prediction required. Type |
| covariance | covariance between test and training individuals. if there are q testing individuals and N-q training individuals, then this covariance matrix is q x (N-q) |
| ... | additional arguments to pass to predict function |
Value
The object returned depends on type.
Details
s is the new vector at which predictions are requested. If
s is not in the lambda sequence used for fitting the model, the
predict function will use linear interpolation to make predictions. The new
values are interpolated using a fraction of predicted values from both left
and right lambda indices. coef(...) is equivalent to
predict(ggmix_fit, type="coefficients",...). To get individual level
predictions at each value of lambda, you must provide the lambda sequence
to the s argument. You can pass either a ggmix_fit or ggmix_gic object. See
examples for more details.
Examples
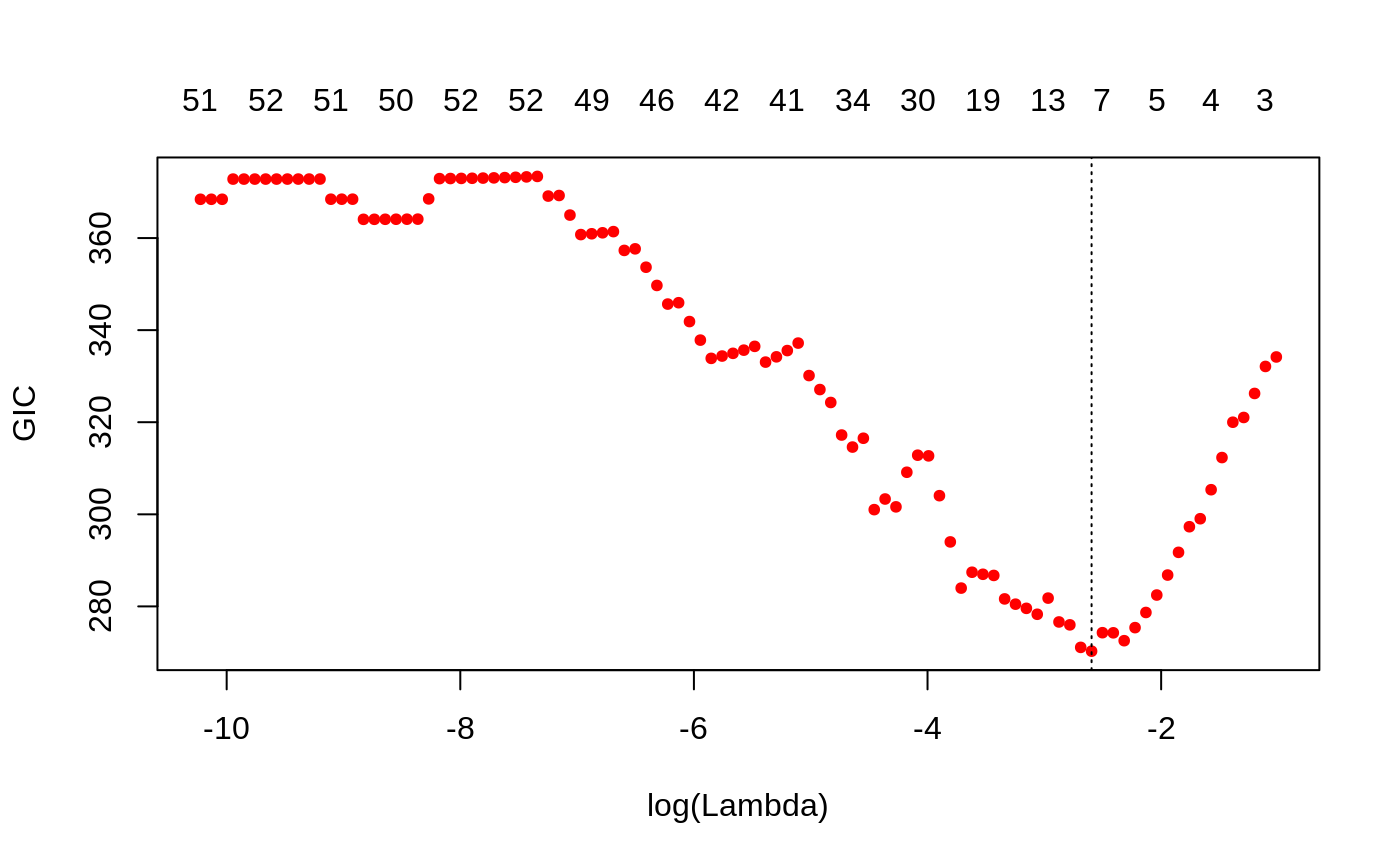
data("admixed") fitlmm <- ggmix(x = admixed$xtrain, y = admixed$ytrain, kinship = admixed$kin_train, estimation = "full") bicGGMIX <- gic(fitlmm, an = log(length(admixed$ytrain))) plot(bicGGMIX)#> 51 x 1 sparse Matrix of class "dgCMatrix" #> 1 #> (Intercept) 0.005271703 #> X23 . #> X36 . #> X38 . #> X40 . #> X53 . #> X74 . #> X83 . #> X114 . #> X139 . #> X143 . #> X168 -0.086106135 #> X176 . #> X244 . #> X246 . #> X249 . #> X266 . #> X271 . #> X273 . #> X282 . #> X286 . #> X300 . #> X302 -0.194013522 #> X312 . #> X315 . #> X330 . #> X336 . #> X344 . #> X348 . #> X352 . #> X375 . #> X383 . #> X403 . #> X404 . #> X420 . #> X422 . #> X431 . #> X435 . #> X441 . #> X447 . #> X468 . #> X485 . #> X503 -0.134716832 #> X512 . #> X515 . #> X516 . #> X407 . #> X507 . #> X524 1.476145530 #> X538 -0.826482104 #> X243 .yhat_test <- predict(bicGGMIX, s="lambda.min", newx = admixed$xtest, type = "individual", covariance = admixed$kin_test_train) cor(yhat_test, admixed$ytest)#> [1] 0.9135335